Projects
1) The Impact of Temperature on the Evolution of Marine Diatoms
Collaborating Project
Katrin Schmidt, Cock van Oosterhout, Taoyang Wu, Andrew Toseland, Matt Clark, Thomas Mock
Recent data on experimental evolution with eukaryotic phytoplankton have revealed that these organisms are capable of responding quickly to environmental change most likely caused by evolutionary processes other than mutations of their DNA (e.g. transposon activation, epigenetics, short non-coding RNAs). Thus, it seems that short-term acclimatory processes as part of their plastic response become accommodated very quickly and therefore significantly contribute to the evolution of these organisms. Information and insights from this project will provide first experimental evidence on how fast evolutionary processes such as epigenetics of key marine primary producers (diatoms) under global warming will impact phenotypes underpinning food web structures and biogeochemical cycles in the future ocean.

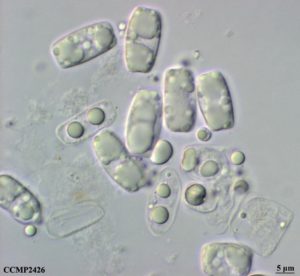
Figure 1: Growth assays with evolved cell lines and the ancestral cell line (T0) of Tp (Unpublished)
So far, there are only preliminary studies (e.g. Kirkham et al. 2017, Schaum et al. 2018) addressing the role of temperature for the evolution of diatom genomes and transcriptomes despite the urgent need to predict the impact of global warming on the evolution and adaptation of key ecological species such as diatoms. To address this question, about 6 years ago, we have initiated a long-term study on the evolution of two key marine diatom species to low and high temperatures. These are the temperate species Thalassiosira pseudonana (Tp) (CCMP1335) and the polar psychrophilic species Fragilariopsis cylindrus (Fc) (CCMP1102). In order to provide the material for our molecular biological and physiological investigations, we have already invested 2.5 years (e.g. 350 generations for T. pseudonana) of effort in cultivating five different biological replicates (in addition to technical replicates) per temperature for each species. First growth assays and preliminary physiological studies were done at T0 and T350 to obtain first insights into the mechanisms of evolution (Figure 1). Currently, we re-sequence 48 cell lines, and we already have obtained transcriptome data for all of them. Furthermore, bisulfite sequencing will reveal the role of DNA methylation for acclimation and adaptation of diatoms to low and high temperatures. Preliminary data from comprehensive transcriptome analyses have revealed that transposable elements seem to play a key role for the adaptation to temperature, and we are curious to see how this is linked with DNA methylation. Furthermore, we will dissect within-population diversity using single-cell genomics.
2) Sea of Change: Eukaryotic Phytoplankton Communities in the Arctic and Atlantic Ocean
Collaborating Project
Kara Martin, Katrin Schmidt, Andrew Toseland, Kerrie Barry, Erika Lindquist, Susannah Tringe, Willem H. van de Poll, Klaas Timmermans, Michael Ginzburg, Ahmed Moustafa, Klaus Valentin, Vincent Moulton, Richard Leggett, Igor V. Grigoriev, Thomas Mock
This JGI (US) and NERC (UK) funded project investigates eukaryotic phytoplankton communities across latitudinal temperature zones of the Arctic and Atlantic Ocean to identify drivers of microalgae biodiversity and biogeography. Temperature is a main driving factor for species distribution on land and in the ocean. Due to global warming, ocean temperatures are rising, allowing temperate phytoplankton species expand their range further north into Polar Regions. This invasion and the sea ice retreat places polar communities at extinction risk, impacting the food web and biogeochemical cycles. We are currently lacking fundamental data on phylogenetic, functional and metabolic diversity in eukaryotic phytoplankton of the Arctic Ocean but also from temperate regions. These data are needed in order to assess and compare the current state of the different biomes and also to predict future changes in biogeochemical cycles in future changing and warmer ecosystems. This integrative project is based on metagenome and metatranscriptome sequencing of samples collected on ca. 70 stations in the Arctic and Atlantic Ocean (Figure 1). There are additional data available on important environmental variables such as nutrients, light, temperature and carbon dioxide. Currently, we are also establishing algal cultures from many of these stations.
For additional information see:
- School of Environmental Sciences, University of East Anglia, Norwich, UK
- School of Computing Sciences, University of East Anglia, Norwich, UK
- U.S. Department of Energy Joint Genome Institute, Walnut Creek, California, USA
- University of Groningen, Netherlands
- Royal Netherlands Institute for Sea Research, Texel, Netherlands
- Alfred-Wegener-Institute, Helmholtz Centre for Polar and Marine Research, Bremerhaven, Germany
- The American University in Cairo, School of Science and Engineering, Cairo, Egypt

Figure 1: 70 stations in the Arctic and Atlantic Ocean
3) Whole genome expression profiling of the polar diatom Fragilariopsis cylindrus
Collaborating Project
Jan Strauss (EMBL, Hamburg, Germany) and Thomas Mock
Diatoms are unicellular photosynthetic eukaryotes with a silicate cell wall. They often dominate polar marine ecosystems, driving the major biogeochemical cycles in these areas. The obligate psychrophilic diatom Fragilariopsis cylindrus is a keystone species in the Southern Ocean. It thrives both in open waters and sea ice and has become a model for studying eukaryotic microalgal adaptations to polar marine conditions. The aim of this thesis was to identify how the genome of F. cylindrus has evolved to cope with marine environmental conditions of the Southern Ocean. To identify key genes, comparative genomics, high-throughput transcriptome sequencing and reverse genetics were applied. Comparative genomics with the sequenced mesophilic diatoms Phaeodactylum tricornutum and Thalassiosira pseudonana was combined with genome-wide RNA-Seq transcriptome analysis, leading to the discovery a new bacteria-like rhodopsin not present in other sequenced diatoms. The characterisation of a bacteria-like rhodopsin in F. cylindrus was conducted by applying reverse genetics tools.
Research Project Outline
Global significance of light-driven proton pumps in eukaryotic marine phytoplankton.
Sunlight is the ultimate source of energy and a major source of environmental information for almost all living organisms. The efficient capture and use of light is a widespread and exquisitely evolved process represented across all kingdoms of life. The metabolic mode in which organisms convert light energy into chemical energy (ATP and NADPH) for growth is known as phototrophy. Two mechanistically distinct processes are known to power phototrophy. While one mechanism employs chlorophyll-containing photochemical reaction centres, the other employs rhodopsins, membrane-embedded photoreceptors composed of opsin proteins and the light-absorbing pigment retinal. Microbial rhodopsins were first discovered in purple membranes of Archaebacteria in the form of the light-driven proton pump bacteriorhodopsin and shown to generate a proton motive force across cell membranes, ultimately powering the production of ATP for energy-requiring processes in the cell. This process is independent from chlorophyll-based photosynthesis. Surprisingly, putative proton-pumping rhodopsins powering phototrophy in prokaryotes were recently also discovered in eukaryotic marine phytoplankton. However, nothing is known about their function in vivo and the role of a light-driven proton-pump in the presence of a proton gradient-generating chlorophyll-based photosynthetic apparatus remains puzzling and speculative. A better understanding of the role of rhodopsins in marine photosynthetic organisms is of global significance, because bacteria-like rhodopsins were shown to be very abundant in metatranscriptome surveys of eukaryotic marine phytoplankton and seem to be of particular significance in iron-limited areas of the ocean, accounting for 35% of the total ocean surface. Furthermore, recent studies suggest that ocean acidification due to increased dissolution of anthropogenic CO2 may decrease iron availability to phytoplankton, which may alter phytoplankton diversity in the oceans, thus, selecting for those phytoplankton species that have a competitive advantage (e.g. by the presence of bacteria-like rhodopsins) under reduced iron concentrations. Therefore we use molecular genetic approaches to elucidate the function and significance of light-driven rhodopsin proton pumps in eukaryotic marine phytoplankton.
4) The role of bacteria-like proton pumping rhodopsins in eukaryotic and photosynthetic marine phytoplankton
Collaborating Project
Jan Strauss (EMBL, Hamburg, Germany), Chris Bowler, Georg Nagel and Thomas Mock
5) Genome project with heterotrophic marine diatoms
Collaborating Project
Thomas Mock, Ryoma Kamikawa (Kyoto University, Japan) and Sophien Kamoun (TSL, Norwich)
With this new project, we are sequencing several different strains of hetertrophic diatoms to reveal first insights into the evolution and adaptation of diatoms to the heterotrophic and parasitic life style. This project includes comparative analyses to other heterotrophic eukaryotes that have lost the ability to photosynthesise such as oomycetes and heterotrophic plants. Nitzschia alba (image down below) is one of the species we currently sequence (genome and transcriptome).